The incorporation of precision genomics into breast cancer care will improve patient outcomes. Clinically actionable genomic alterations can be found in the germline, including in high‐risk predisposition genes and genes relevant to drug metabolism (pharmacogenomics), as well as in the tumour, including cancer driver genes, tumour mutation burden, and mutation signatures that predict treatment responses. Genomic testing in Australia is currently ad hoc, assessing individual genes or gene panels in either the germline or the tumour. However, the genomic features of breast cancers are quite heterogeneous. In the Q‐IMPROvE (Queensland IMplementation of PRecision Oncology in brEast cancer) study, we assessed the feasibility of paired germline/tumour whole genome sequencing as a comprehensive genomic approach to capturing this heterogeneity and the diverse types of reportable genomic events.1
We initiated Q‐IMPROvE, a pilot prospective study, to establish frameworks for patient recruitment, sample collection, whole genome sequencing, variant analysis, and a molecular tumour board for discussing findings (Box 1). The study was approved by the human research ethics committees of the Royal Brisbane and Women's Hospital (HREC/2019/QRBW/48171) and the University of Queensland (2020000203), and was prospectively registered with the Australian New Zealand Clinical Trials Registry (ACTRN12621001285842; 23 September 2021).
Twenty‐nine women (median age, 49 years; range, 26–69 years) with breast cancer with a high risk of recurrence (large tumours or tumours with high‐grade features) and being managed with neo‐adjuvant chemotherapy consented to participation in the pilot study. Paired germline/tumour whole genome sequencing (28 women) was undertaken at the same time as patient chemotherapy sessions, and a curated variant report subsequently discussed by the molecular tumour board. Germline analysis was filtered to include six genes considered clinically actionable in breast cancer (BRCA1, BRCA2, TP53, ATM, CHEK2, PALB2) and pharmacological toxicity polymorphisms. Tumour‐specific genomic alterations, including driver gene mutations, DNA copy number and rearrangements, tumour mutation burden, and mutation signatures were defined. Ten of the women had estrogen receptor (ER)‐positive tumours, eleven HER2‐positive tumours, and seven triple‐negative tumours; complete pathologic response (ie, no tumour remaining after chemotherapy), had been achieved in fourteen women, while fourteen had residual tumours.
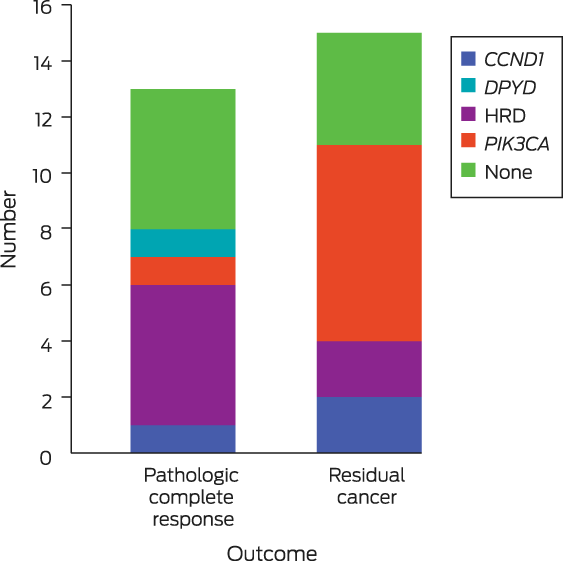
We identified 28 clinically actionable genomic features in nineteen women (germline variants and somatic alterations) (Box 2). Germline variants of clinical importance (BRCA1, CHEK2) were identified in two women who did not meet clinical guidelines for germline genetic testing (ie, estimated risk of finding a pathogenic variant is less than 10%2). Identification of these variants would affect their treatment and risk management in their family. Further, one patient had a DPYD variant (rs55886062) that predicts a cardiotoxic response to 5‐fluorouracil‐based chemotherapy. Given the clinical significance of these findings, these three women were referred for National Association of Testing Authorities (NATA)‐accredited germline DNA testing for confirmation.
Somatically, TP53 (thirteen mutations) and PIK3CA (twelve) were the breast cancer driver genes with the most tumour mutations (as expected); the United States Food and Drug Administration (FDA) has recognised that PIK3CA mutations predict clinical response to alpelisib.3 HER2/ERBB2 were amplified in ten women. A homologous recombination DNA repair defect (HRD) was detected in seven women using HRDetect4 and HRD5 scores, thereby identifying a subset of patients possibly poly(ADP‐ribose) polymerase inhibitor (PARPi)‐sensitive. HRD was linked with a recorded germline alteration in four cases and a mono‐allelic somatic BRCA1 mutation (second/first hit unknown) in one case; for two cases no cause was known.
In conclusion, we found that clinically meaningful outcomes are achievable with matched germline/tumour whole genome sequencing for people with breast cancer, especially women not meeting the criteria for referral for genetic health tests. We have established the logistics and clinical framework for whole genome sequencing as part of breast cancer management in Queensland, and the Medical Research Futures Fund is supporting a national rollout of the program for further evaluation.
Box 1 – The Queensland IMplementation of PRecision Oncology in brEast cancer (Q‐IMPROvE) pilot study: framework*
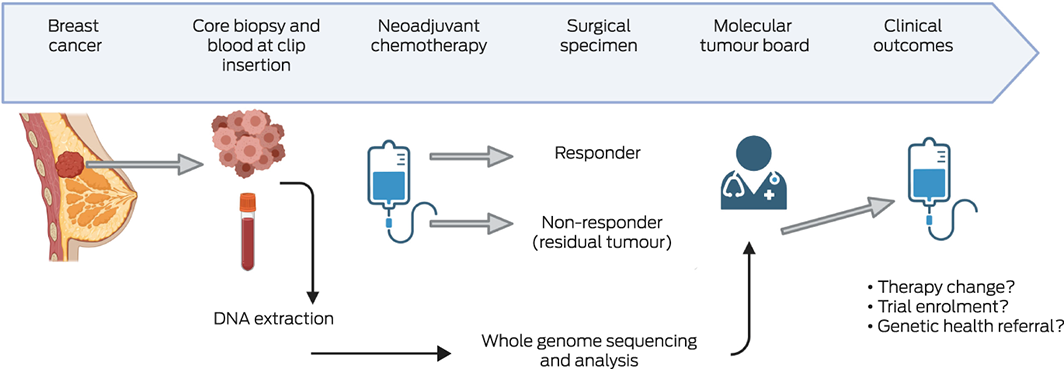
* Following diagnosis of breast cancer and a decision to proceed with neo‐adjuvant therapy, an additional sample (core biopsy) was collected at the time of surgical clip insertion. While the women received chemotherapy, the tissue sample and a pre‐therapy blood sample were sent to Pathology Queensland for DNA extraction and then to BGI Australia (https://bgi‐australia.com.au) for whole genome sequencing. Whole genome sequencing data were analysed by genomiQa, which compiled a comprehensive genomic report on germline variants in a pre‐defined gene list, and somatic alterations in a pre‐defined list of driver genes, tumour mutation burden, mutational signature scores, and homologous recombination deficiency scores (HRD, HRDetect). This report was discussed by the molecular tumour board.
Received 4 October 2022, accepted 7 March 2023
- 1. Vidgen ME, Williamson D, Cutler K, et al. Queensland Genomics: an adaptive approach for integrating genomics into a public healthcare system. NPJ Genom Med 2021; 6: 71.
- 2. Gennari A, Andre F, Barrios CH, et al; ESMO Guidelines Committee. ESMO Clinical Practice Guideline for the diagnosis, staging and treatment of patients with metastatic breast cancer. Ann Oncol 2021; 32: 1475‐1495.
- 3. André F, Ciruelos EM, Juric D, et al. Alpelisib plus fulvestrant for PIK3CA‐mutated, hormone receptor‐positive, human epidermal growth factor receptor‐2‐negative advanced breast cancer: final overall survival results from SOLAR‐1. Ann Oncol 2021; 32: 208‐217.
- 4. Davies H, Glodzik D, Morganella S, et al. HRDetect is a predictor of BRCA1 and BRCA2 deficiency based on mutational signatures. Nat Med 2017; 23: 517‐525.
- 5. Telli ML, Timms KM, Reid J, et al., Homologous recombination deficiency score predicts response to platinum‐containing neoadjuvant chemotherapy in patients with triple‐negative breast cancer. Clin Cancer Res 2016; 22: 3764‐3773.






Correspondence: s.lakhani@uq.edu.au
Open access
Open access publishing facilitated by The University of Queensland, as part of the Wiley ‐ The University of Queensland agreement via the Council of Australian University Librarians.
The study was funded by Queensland Health through Queensland Genomics as an Innovation study. We also acknowledge funding from the Medical Research Futures Fund, Genomics Health Futures Mission.
We thank the participating women and their families. We thank also the Royal Brisbane and Women's Hospital (RBWH), Princess Alexandra Hospital (PAH), and Mater Hospital staff members who helped facilitate the study, as well as Pathology Queensland (PQ), genomiQa, BGI Australia, the University of Queensland (UQ), the QIMR Berghofer Medical Research Institute (QIMRB), Genetic Health Queensland (GHQ), Mater Pathology, and Icon Cancer Care.
The Q‐IMPROvE study group is comprised of Amy E McCart Reed (UQ), Therese McCurry (RBWH), Georgina Hollway (genomiQa), Haidar Al‐Saig (RBWH), Vladimir Andelkovic (PAH), Katharine Cuff (PAH), Margaret Cummings (PQ), David Fairbairn (PQ), Po‐ling Inglis (RBWH), Gillian Jagger (PAH), Helene Johanson (PQ), Lauren Kalinowski (UQ), Olga Kondrashova (QIMRB), Lambros T Koufariotis (QIMRB), Anna Kuchel (RBWH), Rahul Ladwa (PAH), Chiyan Lau (PQ), Ben Lundie (PQ), Helen Mar Fan (GHQ), Nicole McCarthy (Icon), Kathryn Middleton (PAH), Kowsalya Murugappan (RBWH), Mark Nalder (RBWH), Colleen Niland (UQ), Michelle K Nottage (RBWH), Kenneth J O'Byrne (PAH), John V Pearson (QIMRB), Kate Roberts (PAH), Gorane Santamaria Hormaechea (PAH), Cameron Snell (Mater Pathology), Karin Steinke (RBWH), Aneta Suder (RBWH), Diana Tam (RBWH), Euan Walpole (PAH), Natasha Woodward (Mater Hospital), Clement Wong (RBWH), Ho Yi Wong (UQ), Wen Xu (PAH), Peter T Simpson (UQ), Nicola Waddell (QIMRB), and Sunil R Lakhani (UQ, PQ).
Georgina Hollway is the clinical genomics lead at genomiQa.