Based on historical descriptions of clinical symptoms, the extent and rapidity of spread, and associated mortality in the elderly, there is little doubt that epidemic influenza has afflicted humans throughout recorded history, and that, on occasions, severe, widespread, global pandemic outbreaks of influenza have occurred. Our knowledge of the causative viruses of human influenza commenced in 1933 with the infection of an experimental animal — the ferret. This was followed by adaptation of the virus to the laboratory mouse and propagation in the allantois of developing chicken embryos,1 which provided the basis for the production of the first crude, but effective, inactivated influenza virus vaccines2 and, together with the observation that the viruses agglutinate erythrocytes, allowed the initiation of the first in-vitro antigenic analysis of influenza isolates.3
Shortly thereafter, the amazing capacity of influenza viruses to undergo antigenic change was recognised, both through in-vitro antigenic analyses and in a practical sense with the first reported vaccine failures.4 This, together with memories of the devastating “Spanish flu” that had swept the world almost three decades earlier, prompted the formation of a World Health Organization surveillance network for influenza, which has developed into the WHO Global Influenza Programme.5
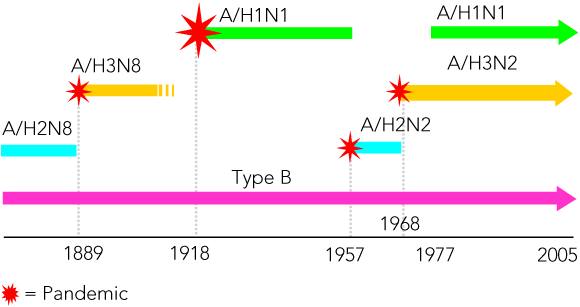
Influenza is a seasonal epidemic disease occurring during the colder months in temperate climates, sometimes accompanied by excess mortality in older adults. In tropical and subtropical countries, infections occur throughout the year, often with one or two peaks of increased activity. Because of this year-round occurrence, the impact of the disease in the tropics is less obvious and only recently have studies shown that it is similar to that in temperate regions.6 Occasionally, and at irregular intervals, severe worldwide pandemic outbreaks occur; these are less constrained by season. Historical evidence suggests that pandemics have occurred at 10–40-year intervals since the 16th century, originating mainly in Asia (Box 1). Three pandemics occurred in the past century: the Spanish influenza pandemic in 1918–1919, which claimed an estimated 50 million lives, and the Asian (1957) and Hong Kong (1968–1969) pandemics, which each resulted in 1–2 million deaths.9,10
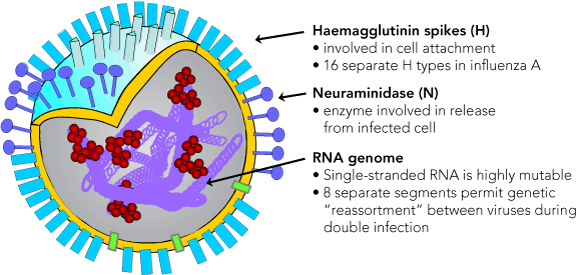
Three types of human influenza viruses have been recognised (types A, B and C), distinguished by their antigenically distinct major internal proteins: the nucleoprotein and matrix protein. These viruses are classified as members of the genus Orthomyxovirus in the family Orthomyxoviridae, and are named according to their type, location where isolated, the successive isolate number from that location, and year of isolation. The virus subtype may also be included (eg, A/Sydney/5/97[H3N2]). Only influenza A and B cause major outbreaks and severe disease; influenza C is associated with common cold-like illness, principally in children.1 Influenza A and B viruses have two major surface glycoprotein antigens: the principal antigen, haemagglutinin (HA or H), responsible for cell attachment, and an enzymically active neuraminidase (NA or N), which assists virus maturation and release (Box 2). For the virus to be infectious, the viral HA must undergo post-translational cleavage into two peptides, HA1 and HA2, by a strong trypsin-like protease.11
These two surface antigens of both influenza A and B undergo gradual, progressive antigenic variation, referred to as “antigenic drift”, which allows the viruses to escape immunity acquired through infection or vaccination and to cause further outbreaks and epidemics. These changes can also be seen by phylogenetic analysis of the HA and NA gene sequences.1 In addition, influenza A viruses with three antigenically distinct HAs and two distinct NAs have circulated in the human population since laboratory studies commenced, and the influenza A viruses are now divided into subtypes based on their HA proteins.12 The Asian and Hong Kong pandemics were found to involve the emergence of new influenza A subtypes in the human population, so few or no people had any immunity to the HA antigens; this is referred to as “antigenic shift”.13 Separate subtypes of influenza B have not been observed, and it does not appear that influenza B is involved in pandemics (Box 3).
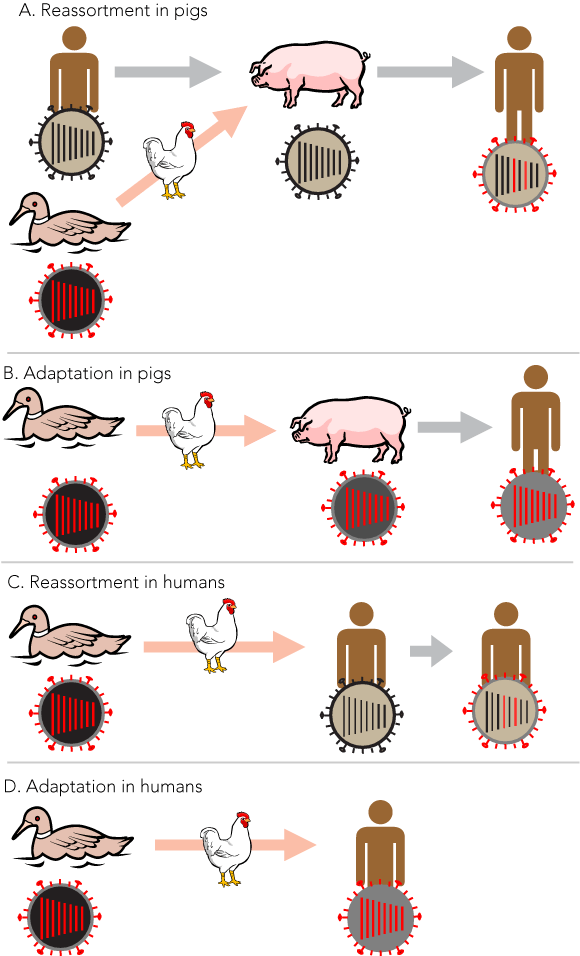
The genomes of influenza A and B viruses consist of eight separate segments of single-stranded, negative-sense RNA. Mutations resulting from the error-prone replication of the single-stranded RNA, and an ability to produce genetic hybrid or reassortant viruses by the mixing of gene segments from two viruses when dual infection occurs both contribute to the evolutionary success of these viruses.11 Although an influenza A virus from swine had been identified in the 1930s, it was not until the 1950s and 1960s that influenza A viruses were found in a number of animal and bird species.14 It was suspected that non-human hosts may play a role in the evolution of pandemic influenza, particularly when it was discovered that viruses with an HA antigenically related to that of the 1968–1969 pandemic virus had been isolated from birds and horses some years earlier. It also appeared that the 1968–1969 pandemic virus may have been derived by genetic reassortment, as the NA antigen was antigenically equivalent to that of the human viruses of the preceding subtype.15 Subsequently, with the benefit of molecular techniques, it was confirmed that the viruses of both the Asian (H2N2) and Hong Kong (H3N2) pandemics were reassortants containing genes derived from the previously circulating human influenza subtype and genes that appeared to have been derived from avian influenza viruses.16
It is now accepted that influenza viruses have evolved in an avian host and that aquatic birds are the major reservoir of influenza A viruses. For influenza A, 16 antigenically distinct HAs and nine NAs have been recognised, all of which can be found circulating in aquatic birds, where they are generally carried asymptomatically.12,17 Some subtypes have been observed to infect and even become established in certain mammals, including horses, swine and humans, and some have been able to infect and cause disease in domestic poultry. Influenza B appears to have diverged and evolved uniquely as a human pathogen some considerable time ago.11 Recently, gene sequences of viruses from the Spanish influenza pandemic were reconstructed from formalin-fixed lung autopsy samples and from unfixed tissue derived from an Alaskan pandemic victim. From these data, it appears that, unlike the 1957 and 1968 pandemic viruses, the 1918 virus may have arisen wholly from an avian influenza virus by adaptive mutations, not by reassortment,18 although this is subject to some debate.19
Influenza viruses display host specificity, and avian viruses do not readily infect humans. A major determinant of this specificity is the difference in cell receptor binding requirements of the HA molecule, and this is determined by differing conformations of carbohydrate (N-acetylneuraminic acid) residues on cell surfaces in different species.20 However, there is evidence that a significant change in receptor preference can occur with as little as a single amino acid mutation at the HA receptor site, and that during evolution of new subtypes in humans, the binding preference for mammalian cell receptors increases with further mutations as a subtype continues to emerge.21 Using technology known as plasmid-based reverse genetics, which allows individual genes to be constructed from known RNA sequences, it has been possible to construct viruses with various genes from a 1918 influenza, and essentially the entire virus, and to study these in the laboratory and experimental animals.22 Contrary to earlier expectations, it appears that no single gene was responsible for the remarkable virulence of this virus — the constellation of all eight gene segments was required to confer full virulence. Because of the distinct host specificity of avian and human influenza, how the adaptive and reassortment events may have occurred remains a matter of conjecture. The domestic pig has been found to carry receptors for both human and avian viruses, and an apparently new disease of swine, more or less coincident with the 1918 pandemic, was found to be due to influenza A/H1N1, which is believed to be closely related to the pandemic strain. Consequently, it has been widely believed that the domestic pig may play a role in adaptation of the viruses to humans and may act as a “mixing vessel” in which reassortment events could occur. This has been demonstrated experimentally.
Although influenza viruses are usually non-pathogenic in their natural waterfowl hosts, two subtypes in particular, H5 and H7, may become highly pathogenic once introduced into domestic poultry. The transition of these subtypes to a highly pathogenic form is associated with acquisition of additional basic amino acids at the cleavage site of the HA molecule. This transforms the virus from one in which cleavage activation of the HA is restricted to the respiratory and intestinal tract, where there are trypsin-like proteases, into one that can be activated by cell-associated proteases that are found throughout the body, resulting in what is referred to as “pantropism”, a much more generalised infection. Historically, outbreaks of the highly pathogenic forms of avian influenza (HPAI) were sporadic, with only 17 episodes reported in the period 1959–1997, and none in Asia.23 With a single exception, the HPAI form of the virus had not been detected in wild aquatic birds and there had been little evidence that humans could be affected by these viruses, with only two recorded cases, both involving H7 virus.24 Nevertheless, serological studies conducted on rural dwellers in southern China had indicated that infection with a number of avian influenza subtypes may not be a rare occurrence.25
Recently, influenza A/H5N1 has developed into an epizootic of domestic poultry with associated human infections, observed first in Hong Kong in 1997, spreading throughout South-East Asia over the following 7 years, and then extending to Russia, Europe, Africa, the Indian subcontinent and the Middle East during the latter part of 2005 and early 2006.26,27 The first indication of this outbreak was a fatal human case of H5N1 infection in Hong Kong in May 1997. This was followed later in the year by a further 17 cases (five fatal), all associated with contact with infected poultry. A cull of poultry in Hong Kong appeared to have controlled this outbreak, but it re-emerged progressively and largely undetected in Asia during 2003 and then caused major outbreaks in Thailand and Vietnam early in 2004, with reports of human cases in which the mortality rate was around 50%. Indeed, by 16 October 2006, the WHO had confirmed 256 cases from 10 countries in Asia and Africa, with 151 deaths — a case fatality rate of 59%.28 Most of the cases have been from China, Indonesia, Thailand and Vietnam.
The reason for the sudden change in the epidemiology of the H5 virus is unclear; a substantial increase in poultry production in Asia, generally under poor housing conditions and often accessible to wild birds, may be a contributing factor. Analysis of the human and avian isolates over the history of these outbreaks has shown that the virus has undergone extensive genetic change, both by repeated reassortment with avian viruses and by mutation, and that a number of antigenic and genetic lineages of virus have emerged in bird populations in different regions (Box 4).29,30 In addition, the highly pathogenic form of the virus has been found widely distributed in aquatic birds, including migratory species, and it appears that these, as well as trade in domestic poultry, have contributed to the widening geographic spread of infection.
Most of the human infections that have occurred appear to have been acquired from direct contact with poultry or poultry products. Although there has been evidence of occasional human-to-human transmission, there is evidence neither of genetic reassortment with human influenza, nor of a significant change of receptor specificity. However, there may be adaptive changes conducive to interspecies transmission,31 and infection of large felids and domestic cats has been reported, with transmission from animal to animal.32 Recent studies indicate that there are receptors for avian influenza in the human lower respiratory tract, which may contribute to the virus occasionally establishing lower respiratory tract infections where it causes severe disease.33 Mortality results from acute respiratory distress syndrome and multiple organ failure, and there is some evidence that this severe pathology is due to dysregulation of the innate immune response with a resulting “cytokine storm”.34 There may also be limited evidence of extra-pulmonary infection with this virus.35,36
Transmission of influenza in humans probably involves both respiratory infection by aerosols and droplets together with some contact transmission from contaminated surfaces.37 Unfortunately, there are insufficient solid data to determine which of these is the more important and to provide guidance in protection of front-line medical staff, particularly regarding the required porosity of facemasks, during a pandemic. Nevertheless, influenza virus is relatively short-lived on most surfaces,38 and clear examples of aerosol transmission have been described in the literature.39 In addition, early experiments indicated greater infectivity by small particle aerosol than by nasal instillation of virus.40
Although cell-mediated immunity plays an important role in recovery from infection, specific immunity to influenza in humans is principally antibody-based. Antibody directed against the HA protects against infection, whereas anti-NA antibody is not protective, but may limit the spread of infection.1 Immunity is type- and subtype-dependent and largely strain-dependent. Consequently, protection against influenza declines as the circulating viruses drift antigenically from the infecting or immunising strains. This necessitates an annual review of vaccine strain formulations, particularly as current vaccines are non-living products targeted to generating humoral immunity against the viral HAs.41 Individuals who have prior exposure to an HA subtype are immunologically “primed” to respond to further viruses of that subtype, which provides some amelioration in the severity of infection and permits vaccination against emerging strains with a single vaccine dose. For unprimed individuals (young children, and, in the event of a pandemic, most or all of the population), infections are likely to be more severe, and successful immunisation usually requires two vaccine doses. A recent re-examination of epidemiological data from the 1957 pandemic suggests that there may be a degree of heterosubtypic (ie, across subtype) immunity acquired through repeated infection, and this may explain the age distribution of infections during a pandemic.42 In addition, coincident with influenza pandemics, the new influenza A virus is observed to totally replace the previously circulating subtype, implying that some form of short-lived heterosubtypic immunity may be involved. These latter two observations may have some relevance to development of more effective influenza vaccines. It is noteworthy that the re-emergence of the H1N1 subtype in the human population in 1977, after an absence of 30 years, did not result in a pandemic and did not replace the circulating H3N2 subtype, presumably because of a level of pre-existing immunity in the population. Consequently, current vaccines have to contain three virus strains, representing the A/H1N1 and A/H3N2 subtypes and type B.
The probability that an avian H5N1 virus from the current epizootic will give rise to a human pandemic cannot realistically be estimated, nor is it possible to foresee the potential severity of any pandemic that might occur. There is laboratory evidence indicating that changes in the receptor binding specificity of H5 could occur, and that there appear to be no genetic barriers to reassortment between H5N1 and contemporary human influenza viruses.43 Some reassortants of this type have recently been made in the laboratory and tested for infection and transmissibility in ferrets, but it appeared that acquisition of human influenza genes alone may be inadequate to create pandemic potential.44 On the other hand, the possibility of adaptive mutation cannot be dismissed and the findings with the 1918 virus suggest that this could possibly result in a virus that is more pathogenic than avian–human recombinant strains (Box 5).
1 Influenza pandemics and probable pandemics during the past three centuries
Compiled from a number of sources, including references 7 and 8. |
|||||||||||||||
4 Evolution of influenza A/H5N1 viruses in Asia
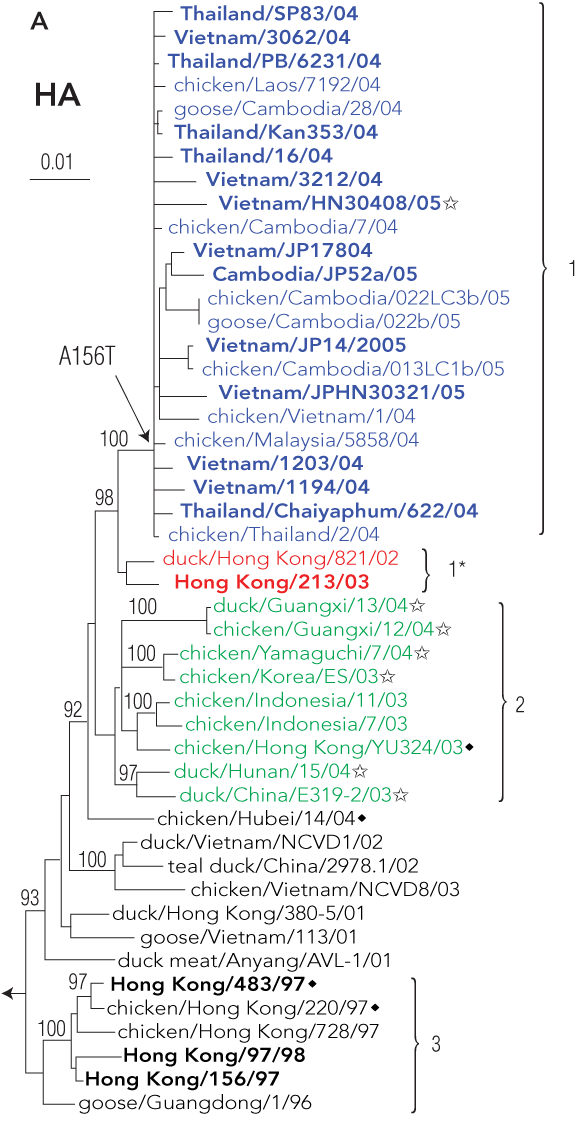
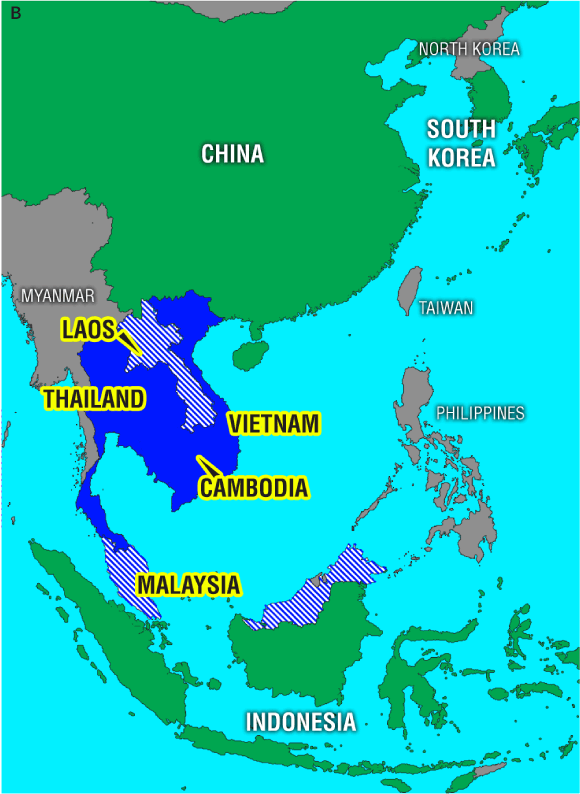
Phylogenetic relationships among H5 haemagglutinin genes from avian influenza A/H5N1 viruses, and their geographic distribution. Viral isolates collected before and during the 2004–2005 outbreak in Asia and selected ancestors are included in the analysis. A: The differing groups or “clades” of haemagglutinin are coloured blue, red, and green. Names in bold denote isolates from human infections. B: Geographic distribution of H5N1 in East Asia: solid blue denotes countries reporting infections with clade 1 H5N1 in humans and birds, cross-hatched blue denotes countries reporting clade 1 H5N1 infections in birds only, and green denotes countries reporting bird infections with clade 2 H5N1. Reproduced from reference 30 with permission from the Centers for Disease Control and Prevention, USA.
- Alan W Hampson1,2
- John S Mackenzie3
- 1 Interflu Pty Ltd, Melbourne, VIC.
- 2 School of Applied Sciences and Engineering, Monash University, Melbourne, VIC.
- 3 Australian Biosecurity Cooperative Research Centre, Curtin University of Technology, Perth, WA.
Alan Hampson holds shares in CSL, and received travel assistance from Roche to attend the Lancet Asia Medical Forum on pandemic influenza, 3–4 May 2006.
- 1. Hampson AW. Influenza virus antigens and ‘antigenic drift’. In: Potter CW, editor. Influenza. Perspectives in medical virology. 7th ed. Amsterdam: Elsevier, 2002: 49-85.
- 2. Wood JM, Williams MS. History of inactivated influenza vaccines. In: Nicholson KG, Webster RG, Hay AJ, editors. Textbook of influenza. Oxford: Blackwell Science, 1998: 317-323.
- 3. Hirst GK. Studies of antigenic differences among strains of influenza A by means of red cell agglutination. J Exp Med 1943; 78: 407-423.
- 4. Francis T Jr, Salk JE, Quilligan JJ Jr. Experience with vaccination against influenza in the spring of 1947. Am J Public Health 1947; 37: 1013-1016.
- 5. Stohr K. Overview of the WHO Global Influenza Programme. Dev Biol (Basel) 2003; 115: 3-8.
- 6. Chow A, Ma S, Ling AE, Chew SK. Influenza-associated deaths in tropical Singapore. Emerg Infect Dis 2006; 12: 114-121.
- 7. Patterson KD. Pandemic influenza 1700–1900: a study in historical epidemiology. Totowa, NJ: Rowman & Littlefield, 1986.
- 8. Potter CW. Chronicle of influenza pandemics. In: Nicholson KG, Webster RG, Hay AJ, editors. Textbook of influenza. Oxford: Blackwell Science, 1998: 3-18.
- 9. Pyle GF. The diffusion of influenza: patterns and paradigms. Totowa, NJ: Rowman & Littlefield, 1986.
- 10. Beveridge WI. The chronicle of influenza epidemics. Hist Philos Life Sci 1991; 13: 223-234.
- 11. Webster RG, Bean WJ, Gorman OT, et al. Evolution and ecology of influenza A viruses. Microbiol Rev 1992; 56: 152-179.
- 12. Fouchier RA, Munster V, Wallensten A, et al. Characterization of a novel influenza A virus hemagglutinin subtype (H16) obtained from black-headed gulls. J Virol 2005; 79: 2814-2822.
- 13. Webster RG, Laver WG. Antigenic variation in influenza virus. Biology and chemistry. Prog Med Virol 1971; 13: 271-338.
- 14. Andrewes CH, Worthington G. Some new or little-known respiratory viruses. Bull World Health Organ 1959; 20: 435-443.
- 15. Webster RG. On the origin of pandemic influenza viruses. Curr Top Microbiol Immunol 1972; 59: 75-105.
- 16. Scholtissek C, Rohde W, Von-Hoyningen V, Rott R. On the origin of the human influenza virus subtypes H2N2 and H3N2. Virology 1978; 87: 13-20.
- 17. Webster RG, Bean WJJ. Evolution and ecology of influenza viruses: interspecies transmission. In: Nicholson KG, Webster RG, Hay AJ, editors. Textbook of influenza. Oxford: Blackwell Science, 1998: 109-119.
- 18. Taubenberger JK, Reid AH, Lourens RM, et al. Characterization of the 1918 influenza virus polymerase genes. Nature 2005; 437: 889-893.
- 19. Gibbs MJ, Gibbs AJ. Molecular virology: was the 1918 pandemic caused by a bird flu? Nature 2006; 440: E8; discussion E9-10.
- 20. Naeve CW, Hinshaw VS, Webster RG. Mutations in the hemagglutinin receptor-binding site can change the biological properties of an influenza virus. J Virol 1984; 51: 567-569.
- 21. Matrosovich MN, Gambaryan AS, Teneberg S, et al. Avian influenza A viruses differ from human viruses by recognition of sialyloligosaccharides and gangliosides and by a higher conservation of the HA receptor-binding site. Virology 1997; 233: 224-234.
- 22. Tumpey TM, Basler CF, Aguilar PV, et al. Characterization of the reconstructed 1918 spanish influenza pandemic virus. Science 2005; 310: 77-80.
- 23. Capua I, Alexander DJ. Avian influenza: recent developments. Avian Pathol 2004; 33: 393-404.
- 24. Swayne DE, Suarez DL. Highly pathogenic avian influenza. Rev Sci Tech 2000; 19: 463-482.
- 25. Shortridge KF. Pandemic influenza: a zoonosis? Semin Respir Infect 1992; 7: 11-25.
- 26. Sims LD, Domenech J, Benigno C, et al. Origin and evolution of highly pathogenic H5N1 avian influenza in Asia. Vet Rec 2005; 157: 159-164.
- 27. World Health Organization. H5N1 influenza timeline. http://www.who.int/csr/disease/avian_influenza/timeline.pdf (accessed Oct 2006).
- 28. World Health Organization. Cumulative number of confirmed human cases of avian influenza A/(H5N1) reported to WHO. 16 October 2006. http://www.who.int/csr/disease/avian_influenza/country/cases_table_2006_10_16/en/index.html (accessed Oct 2006).
- 29. Chen H, Smith GJ, Li KS, et al. Establishment of multiple sublineages of H5N1 influenza virus in Asia: implications for pandemic control. Proc Natl Acad Sci U S A 2006; 103: 2845-2850.
- 30. World Health Organization Global Influenza Program Surveillance Network. Evolution of H5N1 avian influenza viruses in Asia. Emerg Infect Dis 2005; 11: 1515-1521.
- 31. Smith GJ, Naipospos TS, Nguyen TD, et al. Evolution and adaptation of H5N1 influenza virus in avian and human hosts in Indonesia and Vietnam. Virology 2006; 350: 258-268.
- 32. Rimmelzwaan GF, van Riel D, Baars M, et al. Influenza A virus (H5N1) infection in cats causes systemic disease with potential novel routes of virus spread within and between hosts. Am J Pathol 2006; 168: 176-183.
- 33. Shinya K, Ebina M, Yamada S, et al. Avian flu: influenza virus receptors in the human airway. Nature 2006; 440: 435-436.
- 34. Beigel JH, Farrar J, Han AM, et al. Avian influenza A (H5N1) infection in humans. N Engl J Med 2005; 353: 1374-1385.
- 35. Peiris JS, Yu WC, Leung CW, et al. Re-emergence of fatal human influenza A subtype H5N1 disease. Lancet 2004; 363: 617-619.
- 36. Uiprasertkul M, Puthavathana P, Sangsiriwut K, et al. Influenza A H5N1 replication sites in humans. Emerg Infect Dis 2005; 11: 1036-1041.
- 37. Bridges CB, Kuehnert MJ, Hall CB. Transmission of influenza: implications for control in health care settings. Clin Infect Dis 2003; 37: 1094-1101.
- 38. Bean B, Moore BM, Sterner B, et al. Survival of influenza viruses on environmental surfaces. J Infect Dis 1982; 146: 47-51.
- 39. Bell DM. Non-pharmaceutical interventions for pandemic influenza, international measures. Emerg Infect Dis 2006; 12: 81-87.
- 40. Knight V, Gilbert BE, Wilson SZ. Airborne transmission of virus infections. In: Fields B, Martin MA, Kamely D, editors. Banbury report 22: genetically altered viruses and the environment. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory, 1985: 73-94.
- 41. Gerdil C. The annual production cycle for influenza vaccine. Vaccine 2003; 21: 1776-1779.
- 42. Epstein SL. Prior H1N1 influenza infection and susceptibility of Cleveland family study participants during the H2N2 pandemic of 1957: an experiment of nature. J Infect Dis 2006; 193: 49-53.
- 43. Harvey R, Martin AC, Zambon M, Barclay WS. Restrictions to the adaptation of influenza A virus H5 hemagglutinin to the human host. J Virol 2004; 78: 502-507.
- 44. Maines TR, Chen LM, Matsuoka Y, et al. Lack of transmission of H5N1 avian-human reassortant influenza viruses in a ferret model. Proc Natl Acad Sci U S A 2006; 103: 12121-12126.





Abstract
Human epidemic influenza is caused by influenza type A and B viruses, which continually undergo antigenic change in their surface antigens, haemagglutinin (H) and neuraminidase (N).
Influenza epidemics are the consequence of small, ongoing antigenic changes known as “antigenic drift”, which occurs in both influenza types.
Pandemic influenza occurs at irregular and unpredictable intervals, and is the result of a major antigenic change known as “antigenic shift”, which occurs only in influenza A.
Aquatic birds are the evolutionary hosts of influenza viruses; they harbour many distinct forms or subtypes of influenza A, which are usually present as harmless gut infections.
Antigenic shift involves the evolution of a new human influenza A virus through the acquisition of a new haemagglutinin gene encoding a different subtype from an avian influenza, or by the adaptation of an avian virus, causing it to become transmissible between humans.
Two subtypes of avian influenza, H5 and H7, can cause severe infections when introduced into domestic poultry. Recently, influenza A/H5N1 viruses have caused widespread outbreaks, starting in Asia and spreading widely to other regions.
Avian influenza viruses do not readily infect humans. However, during the past 3 years, more than 250 cases of H5N1 infection of humans have occurred, with associated mortality approaching 60%. It is feared that a new pandemic of human influenza may emerge from this.